Mroz pursues COVID-19 clinical clues in DNA code
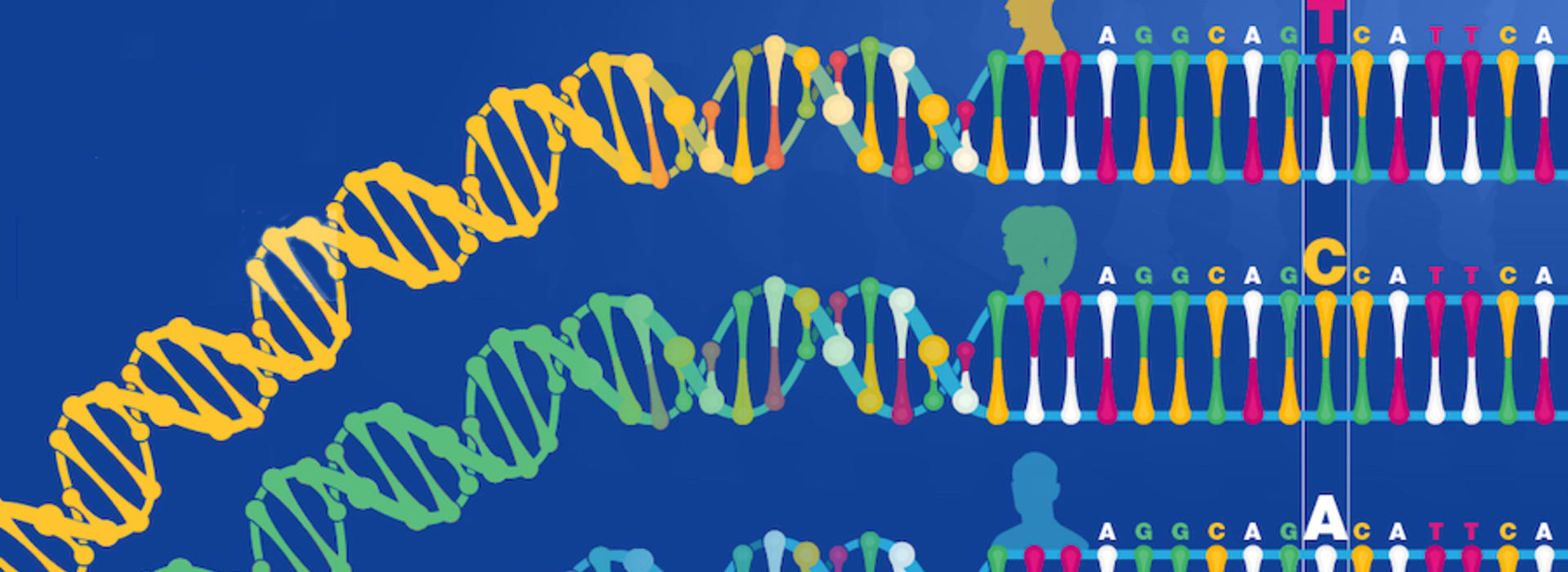
Reading, writing, and editing the human genome, the “book” of human life, is a major scientific enterprise today nearly two decades after the completion of the Human Genome Project was announced.  That enterprise is spawning creative ideas in the COVID-19 era now upon us. LMP assistant professor Pawel Mroz came up with one such idea. Mroz is making a molecular test to detect whether DNA typos called single nucleotide polymorphisms, or SNPs, can yield clues to who is susceptible to SARS-CoV-2 coronavirus infection and which hospitalized patients are at greater risk for severe lung injury and a potentially deadly inflammatory response.
That enterprise is spawning creative ideas in the COVID-19 era now upon us. LMP assistant professor Pawel Mroz came up with one such idea. Mroz is making a molecular test to detect whether DNA typos called single nucleotide polymorphisms, or SNPs, can yield clues to who is susceptible to SARS-CoV-2 coronavirus infection and which hospitalized patients are at greater risk for severe lung injury and a potentially deadly inflammatory response.
“Single nucleotide polymorphism is way too many syllables, so you can understand why we just say ‘snip,’ says Francis Collins, director of the National Institutes of Health who led the Human Genome Project to its completion in 2003. “These are the places in the genome where people differ. In about one out of every 1,000 letters of the code you'll run into one of these where I might have a C and you might have a T, and we'd call that a SNP.”
Collins noted that SNPs typically “don't do very much” because they are usually located in a part of the genome that doesn't have a critical function. But some SNPs do matter, by conferring a higher disease risk or a different drug response -- a rapidly expanding field called pharmacogenomics, which is a major thrust of Mroz’s current research. SNPs have also been implicated as factors in the clinical course of some diseases including immune system response in lung diseases.
COVID-19: "What can we do?"
“When COVID-19 broke out, everyone was thinking ‘What can we do? What’s the mechanism of infection? Can you investigate that?’” Mroz said. He came up with the idea that SNPs in the gene for angiotensin-converting enzyme 2 or ACE2, the receptor in lung epithelial cells by which SARS-CoV-2 enters the body, and SNPs in the gene for TMPRSS2, a protease or protein-cutting enzyme that facilitates viral entry, might have clinical importance. “People have been looking for polymorphisms in ACE2, particularly for ACE inhibitor drugs prescribed for high blood pressure. Polymorphisms of the ACE2 receptor gene have in fact been identified in several different populations. They are associated with differences in how patients respond to ACE inhibitor drug therapy.” Mroz proposes that perhaps these polymorphisms may also affect how people get infected with SARS-CoV-2. “Maybe not every ACE2 receptor has the same affinity for SARS-CoV-2 and maybe not every TMPRSS2 protease primes viral entry into the ACE2 receptor efficiently if polymorphisms are present.”
Mroz took his idea of developing a simple molecular test for ACE2 and TMPRSS2 polymorphisms – what he calls a multiplex SNP-based PCR assay -- to LMP associate professor Bharat Thyagarajan, director of LMP’s Molecular Pathology & Genomics Division. Thyagarajan shared the idea with  Perry Hackett, professor of genetics, cell biology, and development in the College of Biological Sciences, a genome engineering pioneer, and a founder of the Genome Writers Guild, a scientific society involving the University, the Mayo Clinic, Iowa State University, and other institutions. Hackett provided Mroz $25,000 of seed funding for the proposed project through the Hackett Royalty Fund.
Perry Hackett, professor of genetics, cell biology, and development in the College of Biological Sciences, a genome engineering pioneer, and a founder of the Genome Writers Guild, a scientific society involving the University, the Mayo Clinic, Iowa State University, and other institutions. Hackett provided Mroz $25,000 of seed funding for the proposed project through the Hackett Royalty Fund.
Hackett thinks the test Mroz is developing can help address critical aspects of the public health emergency. “In all viral pandemics there are people who may either be resistant to infection due to specific alleles [alternative forms of a gene] that encode viral receptors or people who are asymptomatic due to alternative alleles required for severe disease,” Hackett said. “Identification of people resistant to infection is important for those facing the greatest risk of job-related infection, and identification of asymptomatic carriers is important for lowering risks of further transmission.”
Single DNA letter substitutions influence viral behavior
Mroz said his group is in the early stages of optimizing the assay in LMP’s Molecular Diagnostics Laboratory. “There is data out there that ACE inhibitor drugs may be affected by ACE2 polymorphisms but I don’t think anyone has connected these polymorphisms with patient response, viral load, and other clinical features of COVID-19 patients,” Mroz said. “If we can show that ACE2 receptor gene or TMPRSS2 protease gene polymorphisms affect outcomes, this would be a clinical-grade SNP assay. Individuals that test positive for COVID-19 could then be tested using our multiplex SNP-based PCR assay to predict whether they may need to be followed more closely because they have the separate values associated with higher viral loads and worse outcomes.” Results from his assay, which employs some 25 prominent SNP targets for the ACE2 receptor and TMPRSS2 protease genes, should be available within 24 hours, he said.
Once he has his assay up and running, Mroz said he may form a collaboration with University investigators leading a randomized controlled trial of Losartan, an ACE2 receptor blocking drug, for patients with COVID-19 not requiring hospitalization. The goal of the trial is to see if Losartan can prevent hospitalizations and can prevent lung injury in hospitalized patients with COVID-19 pneumonia. DNA from participant throat swabs would be tested for ACE2 receptor and TMPRSS2 protease SNPs.
 Studies of viral pandemics link SNPs to infectivity, immune response, symptoms, and vaccine effectiveness, demonstrating the power of a single DNA letter substitution to influence viral pathogenesis. Mroz is trying to tap that SNP power through testing in an effort to identify healthy individuals susceptible to infection and infected individuals most at risk for developing severe COVID-19 disease including heart failure and stroke. In the midst of the worst pandemic in a century, select DNA spellchecking may well have something valuable to say about the behavior of a pathogen that has claimed more than 220,000 lives in the U.S. in nine months and more than a million worldwide in less than a year.
Studies of viral pandemics link SNPs to infectivity, immune response, symptoms, and vaccine effectiveness, demonstrating the power of a single DNA letter substitution to influence viral pathogenesis. Mroz is trying to tap that SNP power through testing in an effort to identify healthy individuals susceptible to infection and infected individuals most at risk for developing severe COVID-19 disease including heart failure and stroke. In the midst of the worst pandemic in a century, select DNA spellchecking may well have something valuable to say about the behavior of a pathogen that has claimed more than 220,000 lives in the U.S. in nine months and more than a million worldwide in less than a year.



