Slide Scanning
The University Imaging Centers offers slide scanning services at all three locations.
To submit a request, please fill out the form below.
Click here to submit a slide scanning service order!
Service Overview
Costs for slide scanning services can be found on our rates page.
After we receive your order a UIC staff member will reach out to confirm.
When your scans have been completed we will contact you with instructions for downloading your files. Your physical slides will be returned to the initial drop-off for you to pick up.

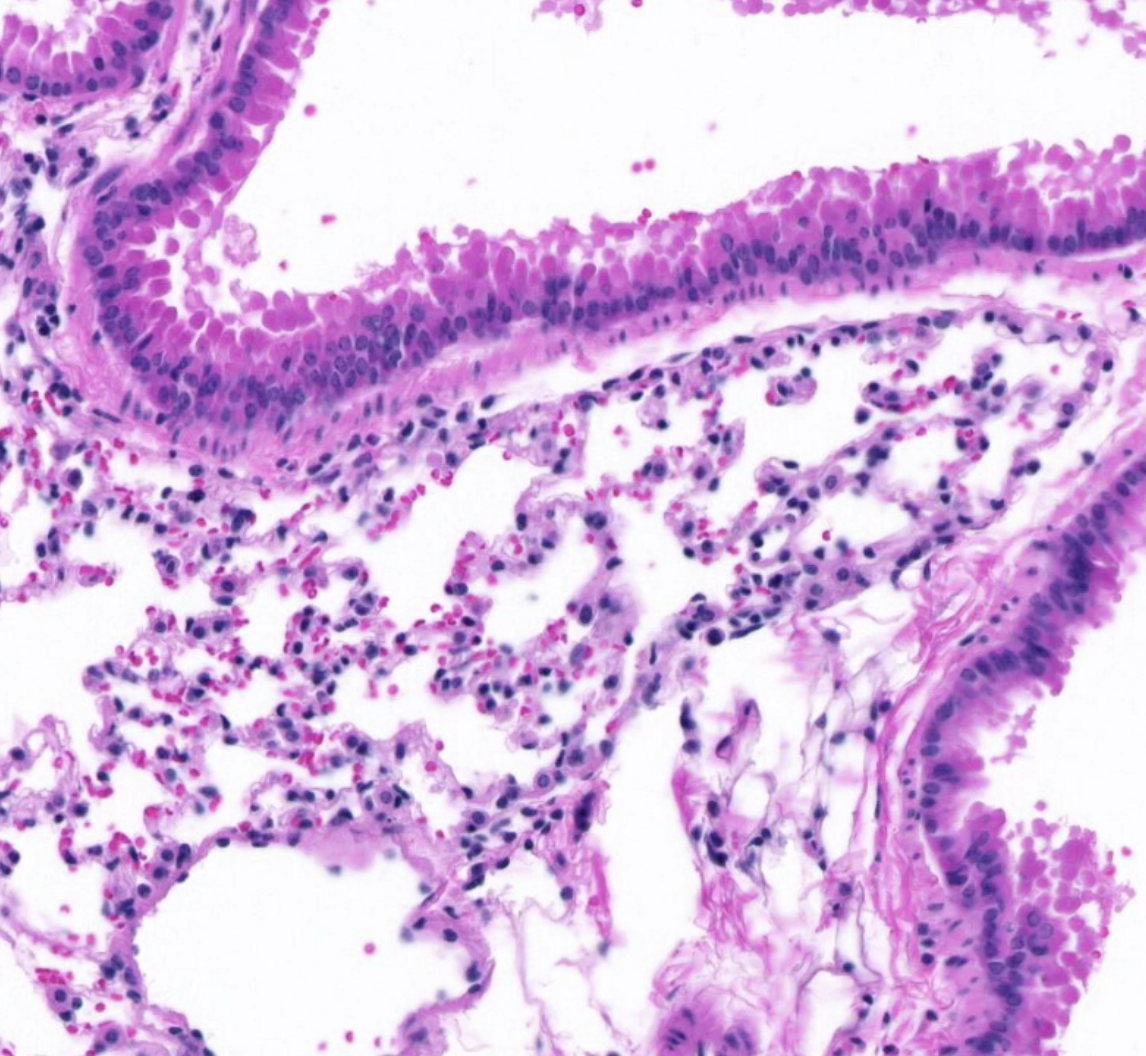
Brightfield Slide Scanning
High-resolution digital Slide Scanning at 20x and 40x (0.75 NA) Brightfield on the Huron TissueScope LE.
Examples:
- Brightfield scan of multiple H&E stained tissues on 1x3" slide at 20X
- Brightfield scan of human tonsil tissue (H&E stained) at 20X
Data File Format:
Scan data is delivered in pyramidal TIFF format via Google Drive or to the provided external device. Huron Viewer and QuPath can be used to open your files.
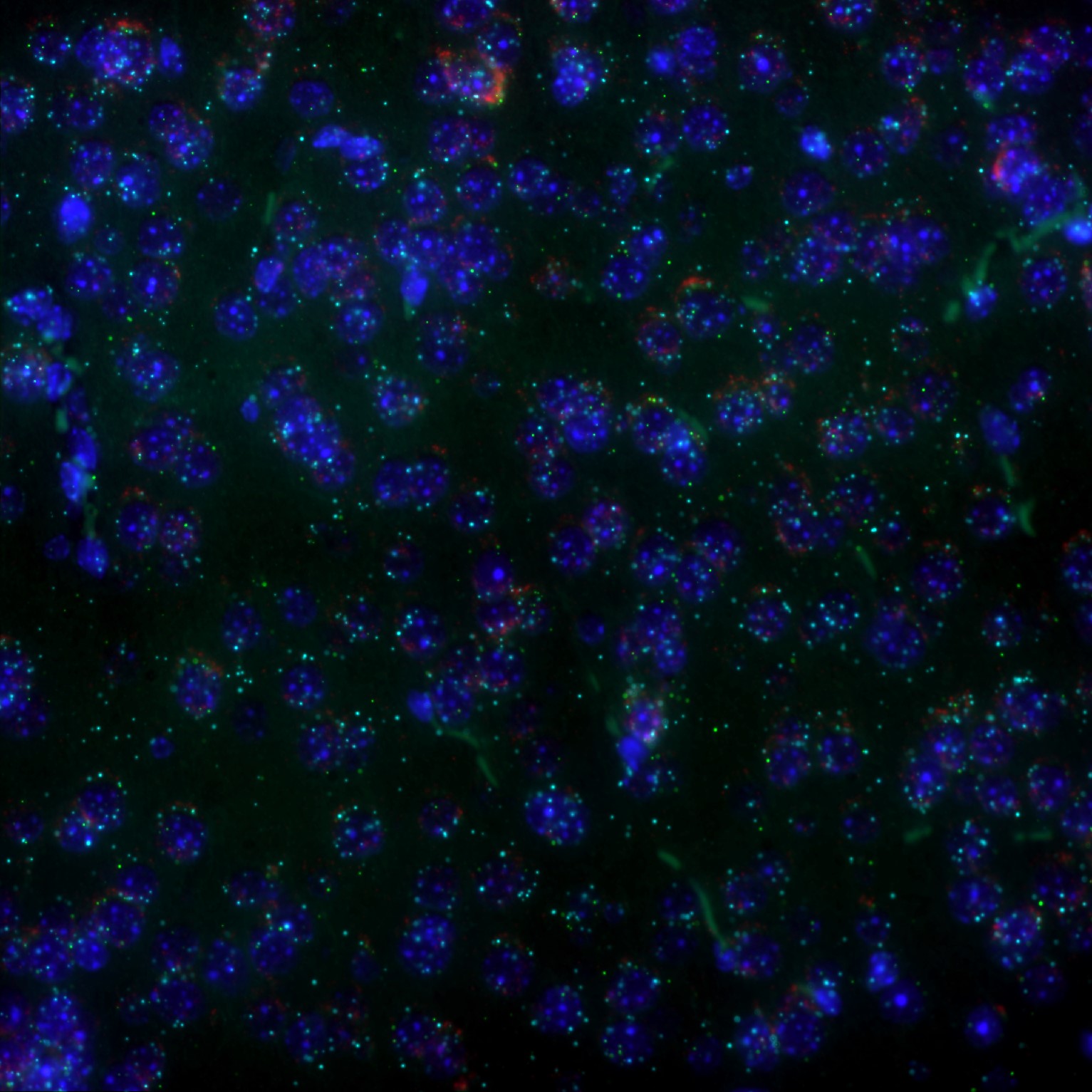
Widefield Fluorescence & RNAscope Slide Scanning
Magnification:
Widefield Fluorescence - 10X or 20X whole slide scans*
*Other magnifications available upon request and may be subject to different rates.
RNAscope - 40X or 60X*
*Optimal magnification and need for Z-stack is determined by UIC Staff based on your region of interest and sample thickness.
Available Channels:
-
DAPI
-
Green - (GFP/AF488/FITC/Opal 520)
-
Orange - (Cy3/AF532/TRITC/Opal 570)
-
Red - (TxRed/Cy5/AF647/AF594/Opal 620)
-
NIR - (Cy7/AF750/Opal 780)
-
Other - (CFP/YFP, etc.)
Data File Format:
Scan data is delivered in the Nikon .nd2 format via Google Drive or to the provided external device. Nikon Elements Viewer and FIJI can be used to open your files.