Zohar Sachs Lab

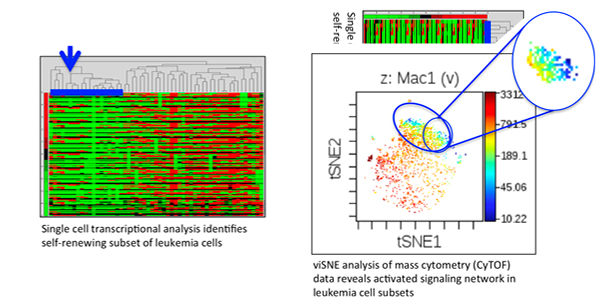
The Sachs Lab's goal is to identify molecular mechanisms of leukemia stem cell self-renewal in acute myeloid leukemia (AML). Since AML cells are highly heterogeneous, we specialize in the application of single-cell, high throughput technologies (including mass cytometry/CyTOF and single-cell RNA sequencing) and patient-derived xenografts and genetically engineered mouse models to address these research questions. We believe that this approach will identify effective therapeutic targets for this deadly disease.
Celebrating cancer 'ResearcHERS' and addressing the challenges
Sachs Lab Members
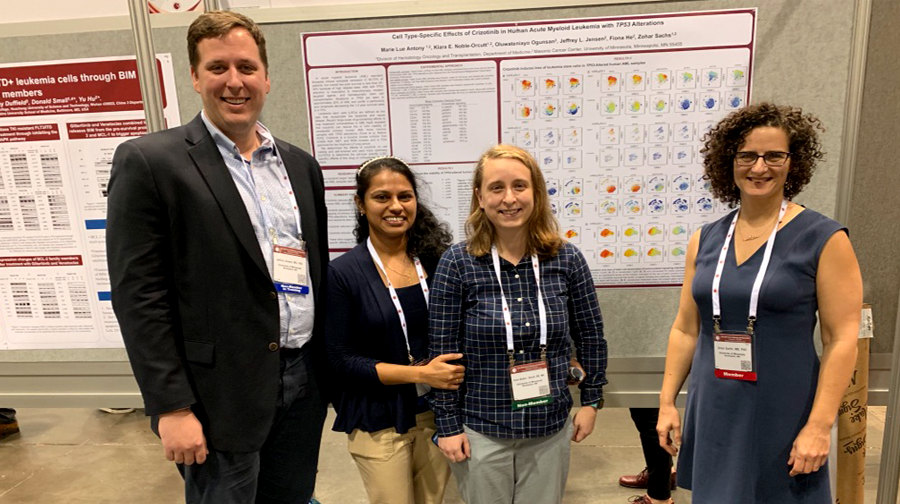

Sachs lab presenting new research findings (“Cell type-specific effects of crizotinib in human acute myeloid leukemia with TP53 alterations”) at the 2019 society for hematology (ASH) annual meeting in Orlando, FL.
Left to right: Jeff Jensen, MD/PhD (former post-doc), Marie Lue Antony, PhD (post-doc), Klara Noble-Orcutt, MS (lab manager), and Zohar Sachs, MD/PhD (principal investigator)

Marie Lue Antony, PhD, Research Associate
Research Interests: Cancer stem cells, AML
Current Research: Therapeutic targeting of leukemia stem cells in AML; understanding the therapeutic vulnerabilities of TP53 altered AML

Yoonkyu (Ryan) Lee, Graduate Student, PhD Candidate, Bioinformatics and computational biology program (BICB)
Research Interests: Cancer, single cell RNA-seq, genomics and machine learning
Current Research: I am developing new computational methods to analyze single cell RNA sequencing data to discover new features and dependencies of self-renewal
Education: BS, Biochemistry, University of Wisconsin (2016)
Email: leex5085@umn.edu

Daniel Chang, Graduate Student, PhD Candidate, Bioinformatics and computational biology program (BICB)
Research Interests: Applying machine learning methods to single-cell RNA sequencing data, cancer stem cell self-renewal mechanisms, and statistical modeling of signaling
Current Research: Using a myelodysplastic syndromes (MDS) mouse model to study the transformation from MDS to secondary acute myeloid leukemia.
Education: BS, Microbiology, University of Minnesota (2013)

Klara Noble-Orcutt, MS, Lab Manager
Research Interests: Cancer stem cells, murine models of leukemia
Current Research: Defining therapeutic vulnerabilities in treatment resistant AML using patient-derived xenografts
Education: BS, MS, University of Minnesota
Email: noble116@umn.edu
Contact
Zohar Sachs, MD, PhD
Assistant Professor
Division of Hematology, Oncology, and Transplantation
Department of Medicine
University of Minnesota
Contact
Sarah Strommer
strom721@umn.edu